By Derek Vargas and Dr. Kushal Suryamohan, MedGenome Inc
Since December 2019, the outbreak of Corona Virus Disease (COVID-19) has posed a serious threat to global health. The number of cases increased quickly and has resulted in over four million deaths worldwide, as of July 2021. In response to this, numerous research projects have been conducted to study the disease etiology, the patterns of epidemic, and potential treatments for the disease. The adaptive immune response plays a central role in clearing viral infections and in turn directly influences patients clinical outcomes. T cells play a crucial role in the immune response to viral infection. Since the start of this pandemic, several innovative tools have been made available for studying the role of T cells in viral infection, and other diseases.
T cells play a critical role in clearing viral infections and providing long-term immune memory. There are two subsets of T cells that participate in the immune response to viral infection. Activated CD8+ T cells directly kill infected cells, while CD4+ T cells produce signaling molecules that drive and support CD8 response and the formation of long-term CD8 memory. CD4+ T cells also participate in the selection and affinity maturation of antigen-specific B cells, which ultimately leads to the generation of neutralizing antibodies. T cells recognize short pathogen-derived peptides presented on the cell surface of the major histocompatibility complex (MHC) using hypervariable T-cell receptors (TCR). TCR repertoire sequencing allows for the quantitative tracking of T-cell clones in patients as these populations go through expansion and contraction. Additionally, TCR repertoire sequencing can provide full length sequences that can be used for biomarker discovery or developing immunotherapies to treat against disease, such as COVID-19.
Despite its widespread adoption, there has been a lack of simple and interactive tools to analyze and explore TCR sequencing data. Many established tools require programming or Unix/Bash knowledge to analyze and visualize results, which can be a barrier for many labs interested in TCR sequencing. To help researchers overcome this hurdle, MedGenome offers end-to-end TCR sequencing services. Using RNA or cells as input, libraries can be generated and sequenced to analyze TCR data, including alpha, beta, delta, and gamma chains. Additionally, we provide a number of analysis outputs, which include critical information such as full length clonotype sequences, V-J usage summaries, CDR3 length distribution, and shared clonotype analysis (Figure 1).
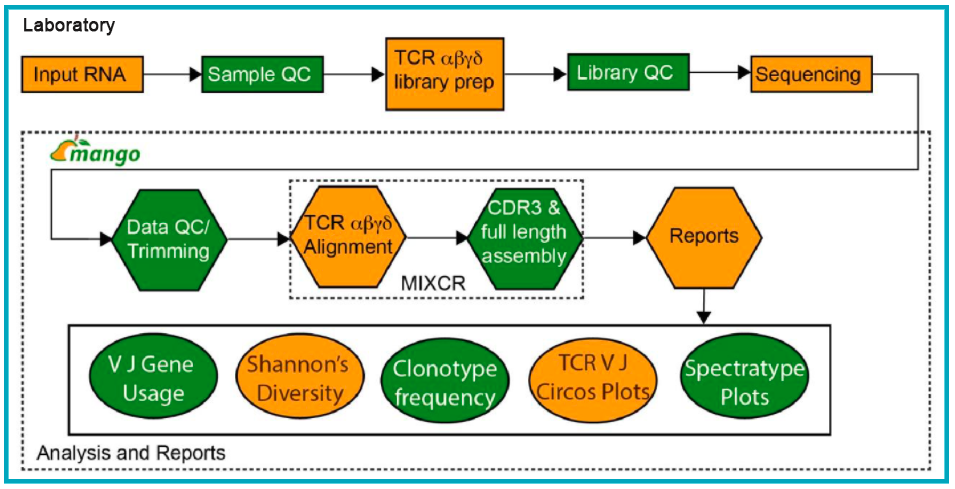
Beyond that, there are additional resources available to scientists interested in studying the immune repertoires of samples. We recently co-hosted a webinar with iReceptor (link below) who discussed the Adaptive Immune Receptor Repertoire Community that they established as a way for researchers to share and access TCR and BCR data. This is a group of immunologists, bioinformaticians, and other experts working together to develop guidelines and standards for the generation, storage, and annotation of this data to facilitate its use by the larger research community. iReceptor facilitates the curation and sharing of TCR seq data from multiple labs and institutions. By engaging this community, scientists can access these datasets, make comparisons with their own data, and effectively increase the amount of data available to answer complex questions about the adaptive immune response.
TCR sequencing data has enormous potential to be used in biomarker and immunotherapy development. The recent pandemic has created a demand for deeper understanding of TCR repertoires. The solutions offered by MedGenome and iReceptor are just a few examples that show how the industry is embracing immunogenomics, and developing solutions to streamline these projects.
References
https://precision.fda.gov/challenges/12/results
https://www.youtube.com/watch?v=JqTHjUY2DlE
#TCR data, #T cells, #TCR sequencing, #immune repertoires, #Adaptive Immune Receptor Repertoire, #TCR and BCR data, #Biomarker, #Immunotherapy
 US
US IN
IN

